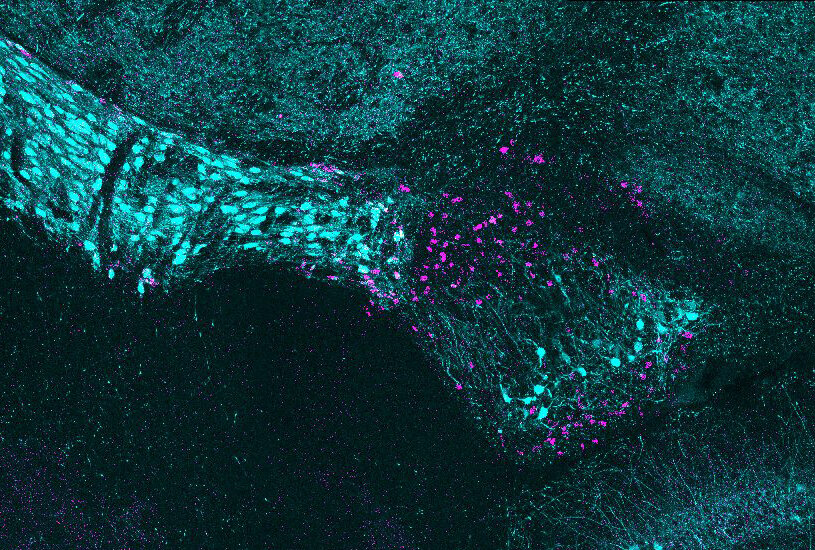

The ventral lateral geniculate nucleus of the brain (vLGN) receives signals from the eye, but is not associated with classical imaging. For decades little was known about the cellular structure and purpose of this region of the brain. In a new study, the Fralin Biomedical Research Institute at VTC Neuroscientists reveals recently identified brain cell subtypes unique in this region that form an amazing layer formation. Credit: Virginia Tech.
Graduate student Ubadah Sabbagh placed a cold glass slide under the microscope. It was getting late and the lab was silent. He adjusted some settings and traced along the optic tract before approaching the thalamus of the brain. What he saw surprised him: two adjacent stripes of bright dots, each dot denoting a cell body.
At that time, two decades had passed since scientists reported that a small area in the brain’s visual thalamus had unique characteristics that set it apart from nearby regions. Previous studies led by other researchers showed that the region, called the ventral lateral geniculate nucleus, connects to the neural circuits involved in the regulation of circadian rhythm and mood.
But little was known about the cellular architecture of the region, until now.
In a new study published online this week in the Neurochemistry journalVirginia Tech scientists identified more than 40 genes expressed in vLGN and discovered more than half a dozen new neuron subtypes, each of which expresses unique molecules and is grouped into very compact layers.
“The ‘amazing’ moment for me was when Ubadah showed me the image of the two cell subtypes aligned in adjacent layers,” said Michael Fox, lead author of the study and a professor at the Fralin Biomedical Research Institute at VTC. “We were surprised because when we see clusters of neurons layered, it generally means that the brain area is secreting different types of information. We now have a more accurate toolkit to help us understand what specific cell types do in vLGN.”
The vLGN spans only a few hundred micrometers in mice and receives signals from the eye through the optic nerve. But unlike other visual regions of the brain, it is not associated with classical imaging.
Neuroscientists initially studied this region of the brain by destroying its cells and documenting the effects. In the process, however, they also disrupted collateral brain circuits. This led to significant behavioral changes and made it difficult to discern what effects were linked to vLGN. Since then, researchers have developed more precise and less invasive genetic tools that allow them to see what happens when the activity of a specific cell type is turned on or off.
Fox wanted to apply these modern techniques to understand the function of vLGN and subsequent connections, but first he needed to determine what types of cells to target. Sabbagh, then a sophomore graduate student in Virginia Tech’s Translational Biology, Medicine and Health graduate program, set out to develop an atlas describing the cellular structure of vLGN as part of his doctoral thesis.
It wasn’t long before they found the first clue that vLGN might be organized by cell type in segregated layers, which the researchers described in a 2018 study.
In that study, Fox’s team revealed two types of specialized lattice-like structures, called perineuronal networks, that surrounded different types of inhibitory neurons. When scientists stained these perineuronal networks, they discovered that different populations of neurons covered by these networks were distributed in different strips across the vLGN.
“That was our first clue that there could be layers, but we still needed to identify more cell types to see if they also aligned in non-overlapping domains,” Fox said.
During the following year, Sabbagh tested approximately 70 different riboprobes to map the vLGN cell landscape. Riboprobes are segments of ribonucleic acids (RNA) that bind to complementary segments of RNA.
Scientists add fluorescence genes taken from other organisms and viruses to the probe, so when an RNA segment meets its corresponding sequence, the target molecules light up. The researchers combined these tools to visualize cells that produce specific genetic molecules in the vLGN, with each molecule glowing a different color under the microscope.
This method revealed different layers in the scans, but the scientists wanted to be sure. Sabbagh coded a computer program to scan vLGN images and measure probe signals, which verified her findings and showed different layers.
But one question persisted: Where were these cells getting their information from?
The researchers used a non-hazardous viral screening tool to determine which neurons in the vLGN communicated with the neurons that line the retina of the eye. The virus took a month to travel a few millimeters between the retina and the thalamus, but it was worth the wait.
The virus revealed that cells in each of the major vLGN layers received direct visual signals from retinal neurons in the eye. Next, collaborators from the University of Louisville analyzed the communication properties of those connections. Together, these ideas help describe how visual information is processed in vLGN.
“This is an exciting finding. We still have more work to do to fill the gaps, but I hope we will find more cell types and perhaps even more layers as we continue to find more biomarkers,” said Fox, who is also a professor at the Virginia Tech College of Science and recently appointed director of the University’s College of Neuroscience.
His team has begun to analyze gene expression in individual cells, using a process called single-cell RNA sequencing. Fox says that once they know more about specific cell types, they can trace the circuits and begin to reveal how vLGN neurons interact with other brain regions.
“We suspect that vLGN has numerous functions, including mood regulation, but now we can look at how specific cell types contribute to those functions more precisely,” said Sabbagh, who recently received a six-year award from the Institutes. National $ 390,000 Health Prize to support your research.
This research also raises questions about brain evolution and comparative biology. As mammals evolved into larger species, their vLGNs decreased relative to the rest of the visual thalamus. By revealing the complicated architecture of vLGN in mice, Fox hopes to shed light on how these genetic instructions might interbreed in other species.
Scientists shed light on striking visual development patterns
Ubadah Sabbagh et al, Various GABAergic neurons organize into subtype-specific sublamines in the ventral lateral geniculate nucleus, Neurochemistry journal (2020). DOI: 10.1111 / jnc.15101
Provided by Virginia Tech
Citation: Scientists discover cellular structure of misunderstood visual brain region (2020, June 24) retrieved on June 25, 2020 from https://medicalxpress.com/news/2020-06-scientists-cellular-poorly- understood-visual.html
This document is subject to copyright. Other than fair dealing for private research or study purposes, no part may be reproduced without written permission. The content is provided for informational purposes only.