
While the Kovid-1p epidemic is such that it was already recognized for the threat it posed, researchers rushed to find anything that could block the spread of the virus. While the vaccine has attracted a lot of attention recently, there was also hope that we could develop a treatment that would block the bad effects of the virus. Most of these are highly practical: identify enzymes that are needed to replicate the virus, and test drugs that block similar enzymes from other viruses. These drugs are designed to be relatively easy to store and administer and in some cases, have already been tested for the safety of humans, so they make reasonable choices for quickly getting something ready for use.
But the devices we have developed in biotechnology allow us to do some less practical things, and a paper released today describes how they can be used to deactivate SARS-COV-2. This is by no means a practical cure, but it does provide a fantastic window into what we can do with biological manipulation.
Throw it in the trash
The whole effort described in the new paper is centered on a simple idea: if you knew how to destroy one of the key proteins of a virus, it would not be able to infect anything. And, easily, our cells have a system of destroying proteins, as it is often useful to do. In some cases, the destroyed protein is damaged; In others, proteins are formed and destroyed at elevated speeds, allowing the cell to respond quickly to changing conditions. In some cases, changes in the environment or activation of signaling pathways can lead to extensive protein destruction, causing the cell to rapidly change its behavior.
This system relies on a small protein called “ubiquitin”. When a protein is to be targeted for destruction, an enzyme called ubiquitin ligase chemically binds a chain of ubiquitin to it. This serves as a toxin recognized by enzymes that digest any protein with the ubiquitin attached to them.
So, the idea behind the new function is to identify the key viral protein, and figure out how to attach it to ubiquitin. This cell will then take care of the rest, digest the viral protein and thus block the production of any useful virus in that cell. In this case, the researchers decided to target the spike proteins that sit on the surface of the coronavirus and allow them to attach and infect new cells.
Unfortunately, there is no protein that binds ubictin to viral spike proteins. Or, rather, there is no protein that corresponds to that description.
But the Harvard team has now built it.
Bioengineering
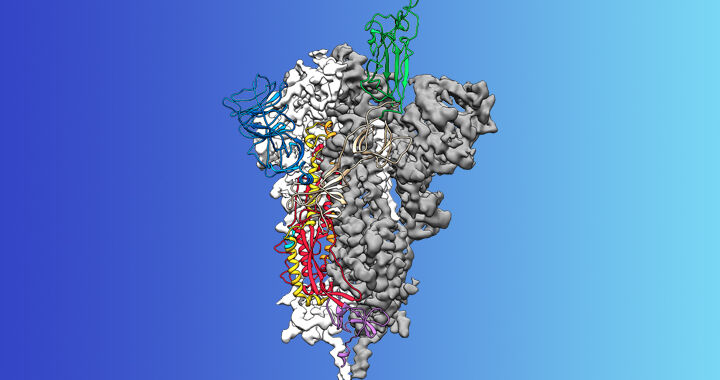
The team’s method of doing so began with the fact that we know something that sticks to the viral spike protein: the cellular protein that pushes it into the cell. It is called angiotensin-converting enzyme 2 or ACE2, but we will call it green protein because in this figure we use color. The idea was to find a part of this protein that stuck to the spike (aka red protein) and bind it to the ubiquitin adding protein (blue). Sounds simple enough.

John Timmer
But there is a complication: green protein also sticks to other proteins found on healthy, unhealthy cells. So, if you are not careful, your virus-destroying enzyme will also destroy the proteins needed for the health of non-protected cells. Which would be a rather large “hey.”
To solve this problem, the researchers downloaded data that showed molecular-level details of the composition of red and green proteins, as well as how those proteins interact. (Yes, it is available.) They then transferred the data to a software software package that detects the most intensively selected interactions between proteins. (Yes, it exists.) They told the program to virtually cut the green protein and find small pieces that satisfied two conditions: the pieces stuck to the red protein of the virus but not the one found on the surface of healthy human cells. .
With the identification of a red-specific bit of green protein, the researchers linked it to something stuck with blue protein, which would bind ubiquitin to red. This hybrid will act as a bridge, binding the viral red protein to the blue dye that binds ubiquitin to it.

John Timmer
This worked, but not particularly well. The authors associate the spike (red one) with a fluorescent protein and found that the fluorescence decreased by about 30 percent compared to the production of their hybrid protein. Better than nothing – but not great.
.Putimization
So, how to make it better? The researchers used a software package to transform each of their green protein fragments, and they examined what the viral spike did to its relationship to the protein. Whatever seemed promising, they engineered real protein. One of these significantly increased the impact; Now, instead of reducing fluorescence by 30 percent, it dropped by 50 percent.
But that was not the end of their efforts. They served as bridges sticking to both the green fragment / linker hybrid, the red protein and the ubiquitous-connective blue. To further the efficiency, the researchers made things a little easier by combining the blue enzyme directly with the green piece. Instead, red has a direct link with the sticking protein and the blue ensuring its destruction. This results in a 60 percent reduction in the amount of fluorescent spike protein present in the cells.
So, a wonderful application of biotechnology, right? Unfortunately, that too is likely to be completely useless, and not because we don’t know if a 60 percent reduction makes sense. For this to be effective, it has to be made by cells because they have an active infection. Which means we have to insert genes that encode the proteins that make up the cells, at least temporarily. We can do that for sure – it’s a technology that depends on some of the leading vaccine candidates. But to get the vaccine to work, we don’t need genes to be active in many cells. To protect an entire organ, we probably.
Conclusion: This is likely to be a non-starter, especially given that there are promising vaccines and many other potential therapies in the pipeline for safety testing. Still, things that make this type of technology wildly impractical for treating humans for the virus cannot be applied to other use cases such as bacteria, crops, animals, or less urgent medical needs. So, while the details of this work are not really important, it is worth noting the fact that we have developed all the underlying techniques required for it.
Communications Biology, 2020. DOI: 10.1038 / s42003-020-01470-7 (About DOI).